Team III Gene Prediction Group: Difference between revisions
| Line 17: | Line 17: | ||
===[https://github.com/tseemann/barrnap/ '''Barrnap''']=== | ===[https://github.com/tseemann/barrnap/ '''Barrnap''']=== | ||
Barrnap predicts the location of ribosomal RNA genes in genomes by using HMMER 3.1 for HMM searching in RNA:DNA style. It supports bacteria (5S,23S,16S), archaea (5S,5.8S,23S,16S), metazoan mitochondria (12S,16S) and eukaryotes (5S,5.8S,28S,18S). | Barrnap predicts the location of ribosomal RNA genes in genomes by using HMMER 3.1 for HMM searching in RNA:DNA style. It supports bacteria (5S,23S,16S), archaea (5S,5.8S,23S,16S), metazoan mitochondria (12S,16S) and eukaryotes (5S,5.8S,28S,18S). | ||
=='''Results: Non-Coding'''== | =='''Results: Non-Coding'''== | ||
Revision as of 19:54, 7 March 2020
Non-Coding homology - All tools initially used
ARAGORN
Aragorn is a computer program identifies tRNA and tmRNA genes. The program employs heuristic algorithms to predict tRNA secondary structure, based on homology with recognized tRNA consensus sequences and ability to form a base-paired cloverleaf. tmRNA genes are identified using a modified version of the BRUCE program.
Infernal
Infernal is for searching DNA sequence databases for RNA structure and sequence similarities. It is an implementation of a special case of profile stochastic context-free grammars called covariance models (CMs). A CM is like a sequence profile, but it scores a combination of sequence consensus and RNA secondary structure consensus, so in many cases, it is more capable of identifying RNA homologs that conserve their secondary structure more than their primary sequence.
Non-Coding Ab initio - All tools initially used
RNAmmer
RNAmmer predicts ribosomal RNA genes in full genome sequences by utilizing two levels of Hidden Markov Models: An initial spotter model searches both strands. The spotter model is constructed from highly conserved loci within a structural alignment of known rRNA sequences.
Barrnap
Barrnap predicts the location of ribosomal RNA genes in genomes by using HMMER 3.1 for HMM searching in RNA:DNA style. It supports bacteria (5S,23S,16S), archaea (5S,5.8S,23S,16S), metazoan mitochondria (12S,16S) and eukaryotes (5S,5.8S,28S,18S).
Results: Non-Coding
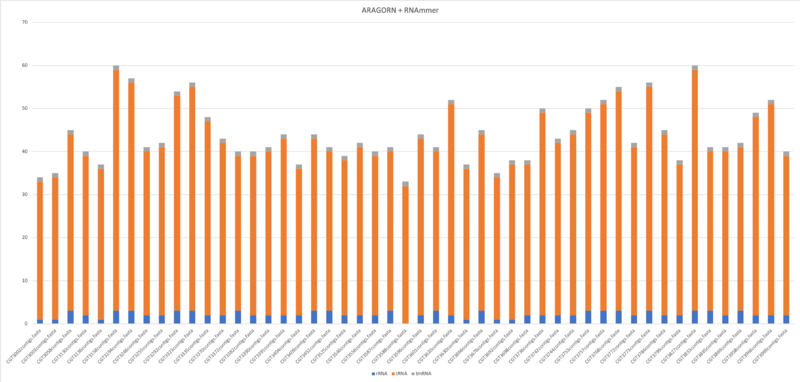
Aragorn & RNAmmer
$ aragorn -l -t -gc1 -w input.fasta -fo –o output.fasta $ aragorn -l -m -gc1 -w input.fasta -fo –o output.fasta
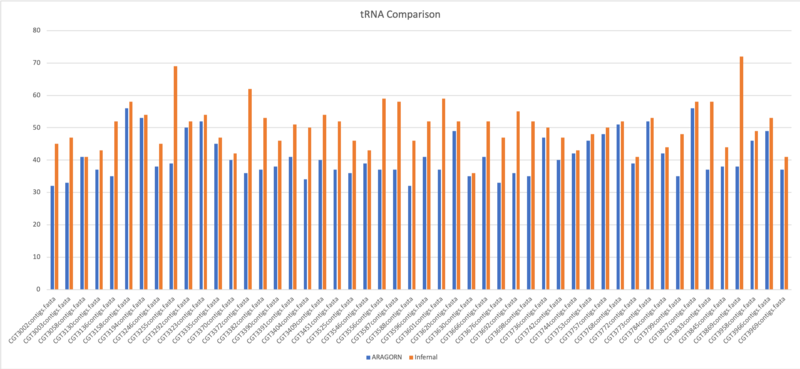
- Average of tRNA: 40.9
- Average of tmRNA: 1
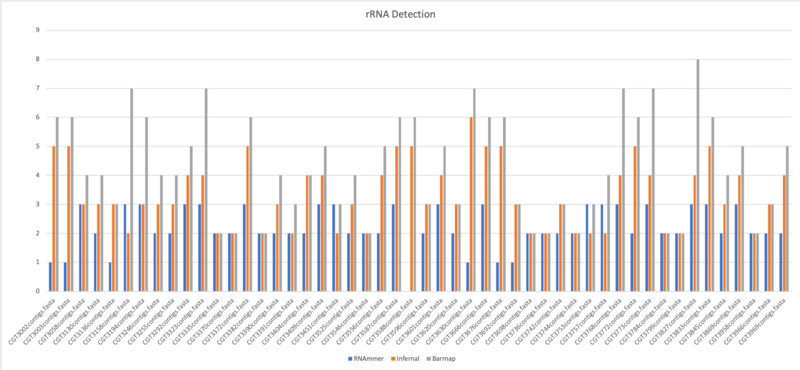
- Average of rRNA: 2.2
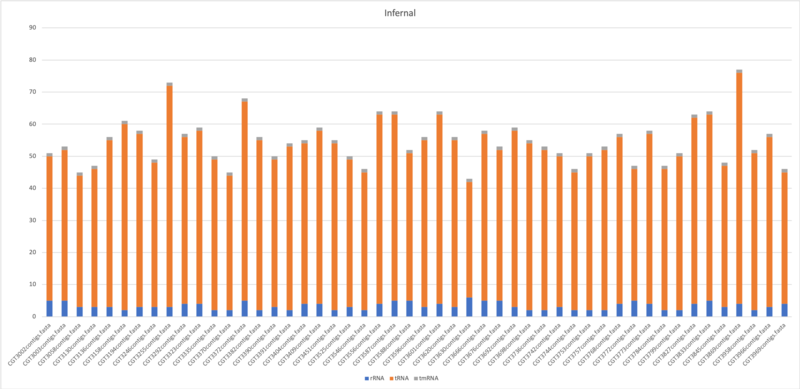
Infernal
$ cmscan --cut_ga --rfam --nohmmonly --tblout $output/$(basename $filename .fasta).tblout --fmt 2 --clanin Rfam.clanin Rfam.cm $filename > $output/$(basename $filename . fasta).cmscan
- Average of tRNA: 50.5
- Average of tmRNA: 1
- Average of rRNA: 3.34